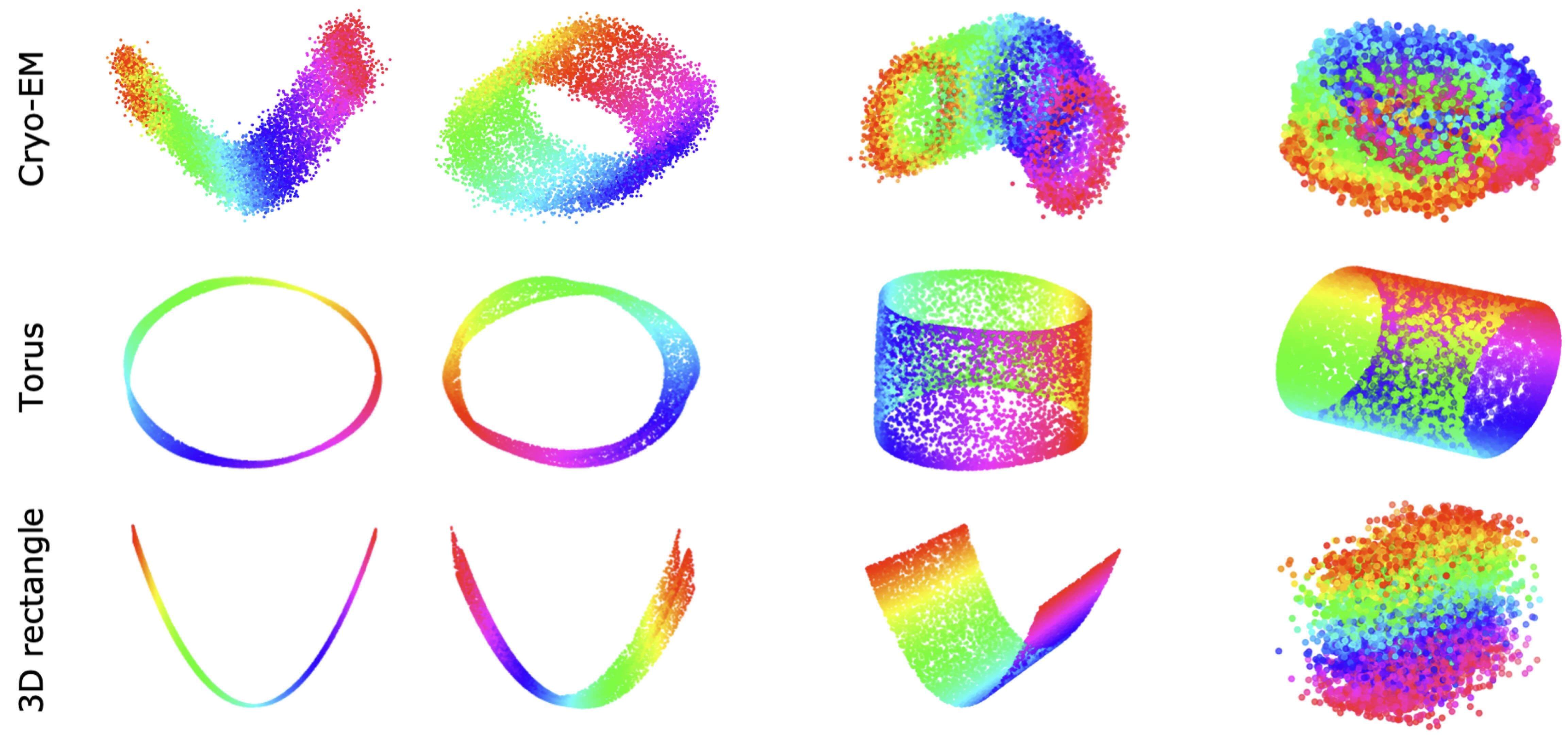About
I'm a senior lecturer (=tenure-track assistant professor) at the department of statistics and operations research, school of mathematical sciences, Tel Aviv university. Prior to that, I was a postdoctoral research associate at Princeton university's program in applied and computational mathematics, working in Amit Singer's group. I did my Ph.D. at the department of computer science and applied mathematics at the Weizmann institute of science, where Boaz Nadler was my doctoral advisor.
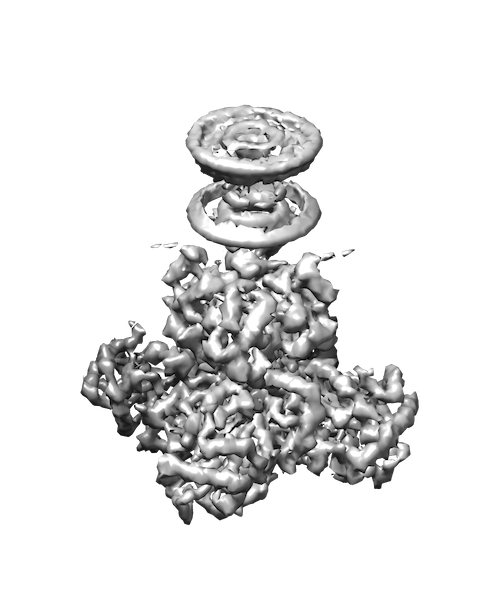
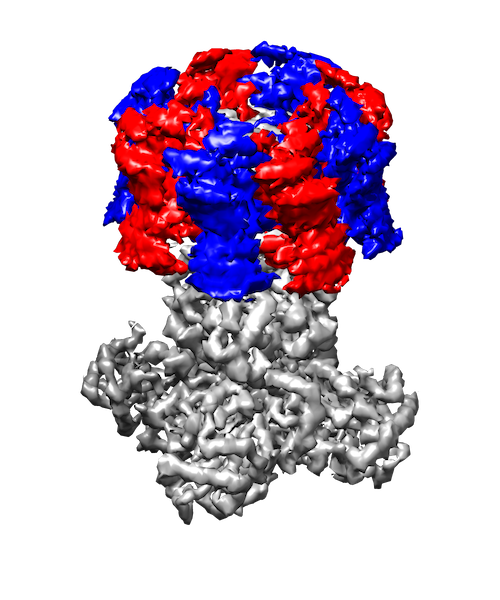
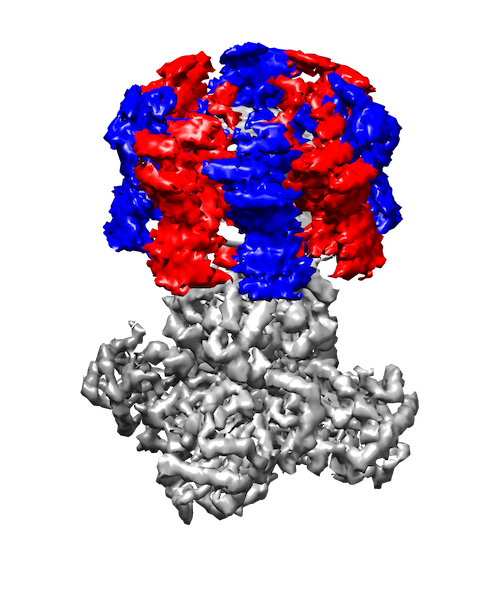
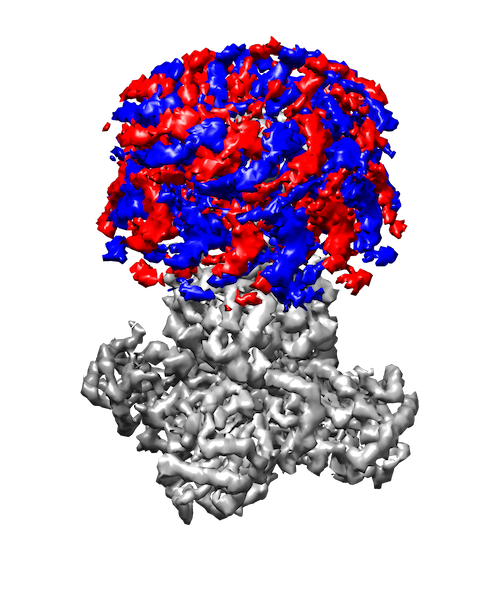
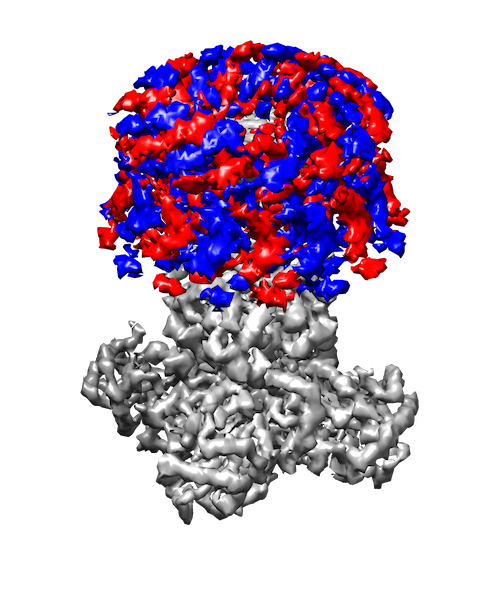
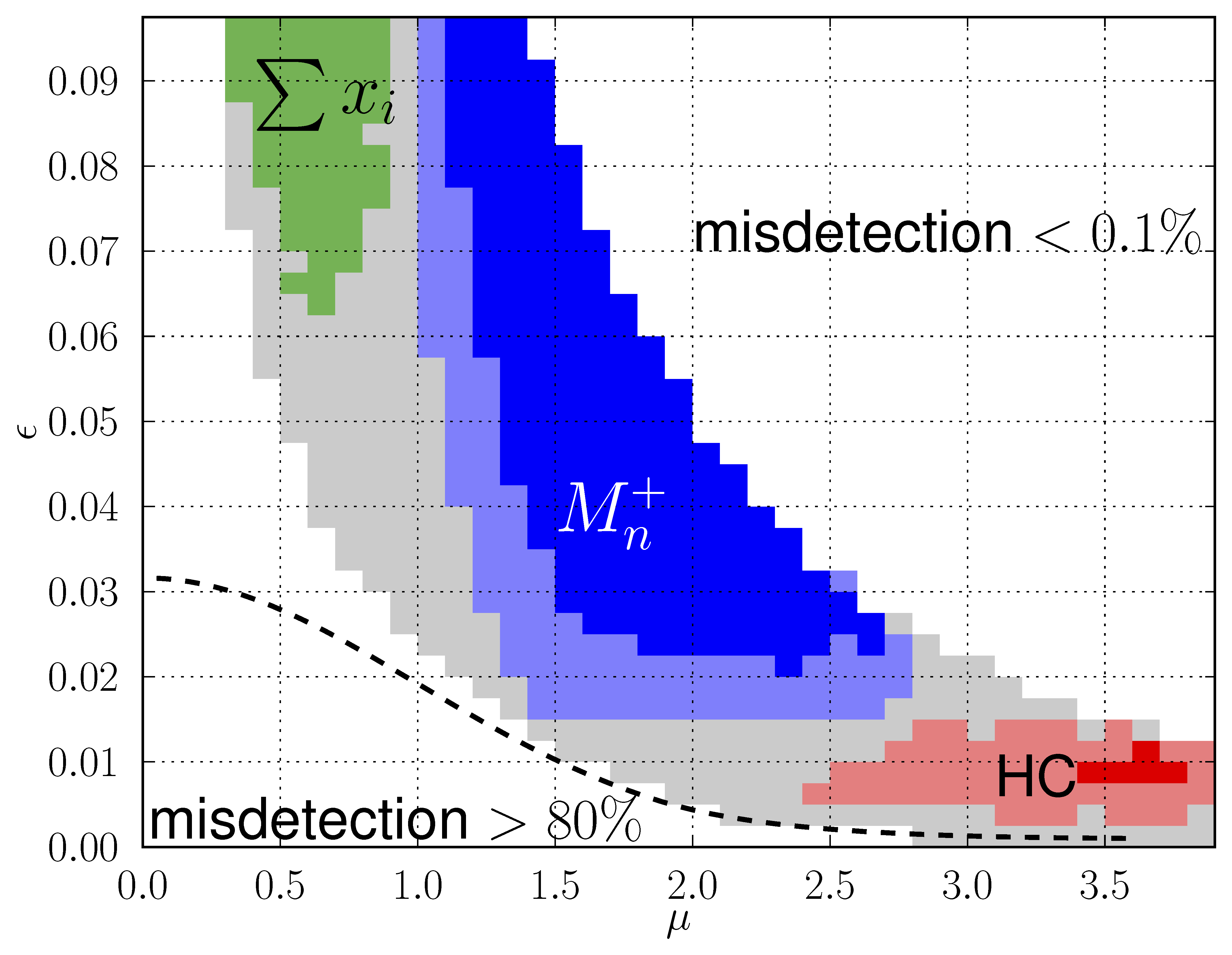
My research interests are broadly in the development of methodology for statistics and machine learning. More specifically, my current focus is developing tools for mapping and analyzing large volumetric data sets. This is motivated by a key challenge in structural biology: the 3D reconstruction and analysis of flexible proteins and other macromolecules from cryo-electron microscopy data sets.
Interested in doing research with me? I sometimes have openings at the MSc/PhD/Postdoc level for people with a strong mathematical foundation. Please email me and we'll setup a meeting.
Interested in statistics/data science research at TAU? If you're looking for an advisor, or research project, or both, or just broadly interested in statistics and data science research, you are welcome to join the TAU Statistics seminar. It normally runs every Tuesday at 10:30-11:30 during the semester, but please check the schedule.
Video Presentations:
• One-World Cryo-EM talk:
Tools for heterogeneity in cryo-EM: manifold learning, disentanglement and optimal transport
• Broad overview of my research (as of 3/2021) at INRIA's DataShape seminar:
Nonparametric estimation of high-dimensional shape spaces with applications to structural biology
• Presentation for 3rd year math students at Tel Aviv university (hebrew):
Shape spaces, dimensionality reduction and product manifold factorization
Teaching
I regularly teach introductory CS and programming classes to students in statistics and data science:
"Introduction to computers for statisticians" and "Advanced programming for data science".
If you are signed up to these classes please use TAU's Moodle to access class recordings and assignments.
I also regularly supervise undergraduate and graduate seminars in statistical learning.
These work-in-progress Lecture notes on Python Programming may be of interest to aspiring data scientists who wish to learn Python programming.
The notes are designed for newcomers and offer an example-based approach with an emphasis on interactive data analysis in the IPython interpreter.
If these notes helped you learn programming please email me!
Funding:
• Israel Science Foundation (ISF)
• United States-Israel Binational Science Foundation (BSF)
• United States National Science Foundation (NSF)



Address: Schreiber 202, School of Mathematical Sciences, Tel Aviv University.